In current methods, determination of the deconvolution model best supported by the data is done manually through comparison of residual error values, which can be time consuming and requires model selection by the user. 
The method also allows for fitting of intermediate exchange spectra, which is not supported by current software in the absence of a specific kinetic model. 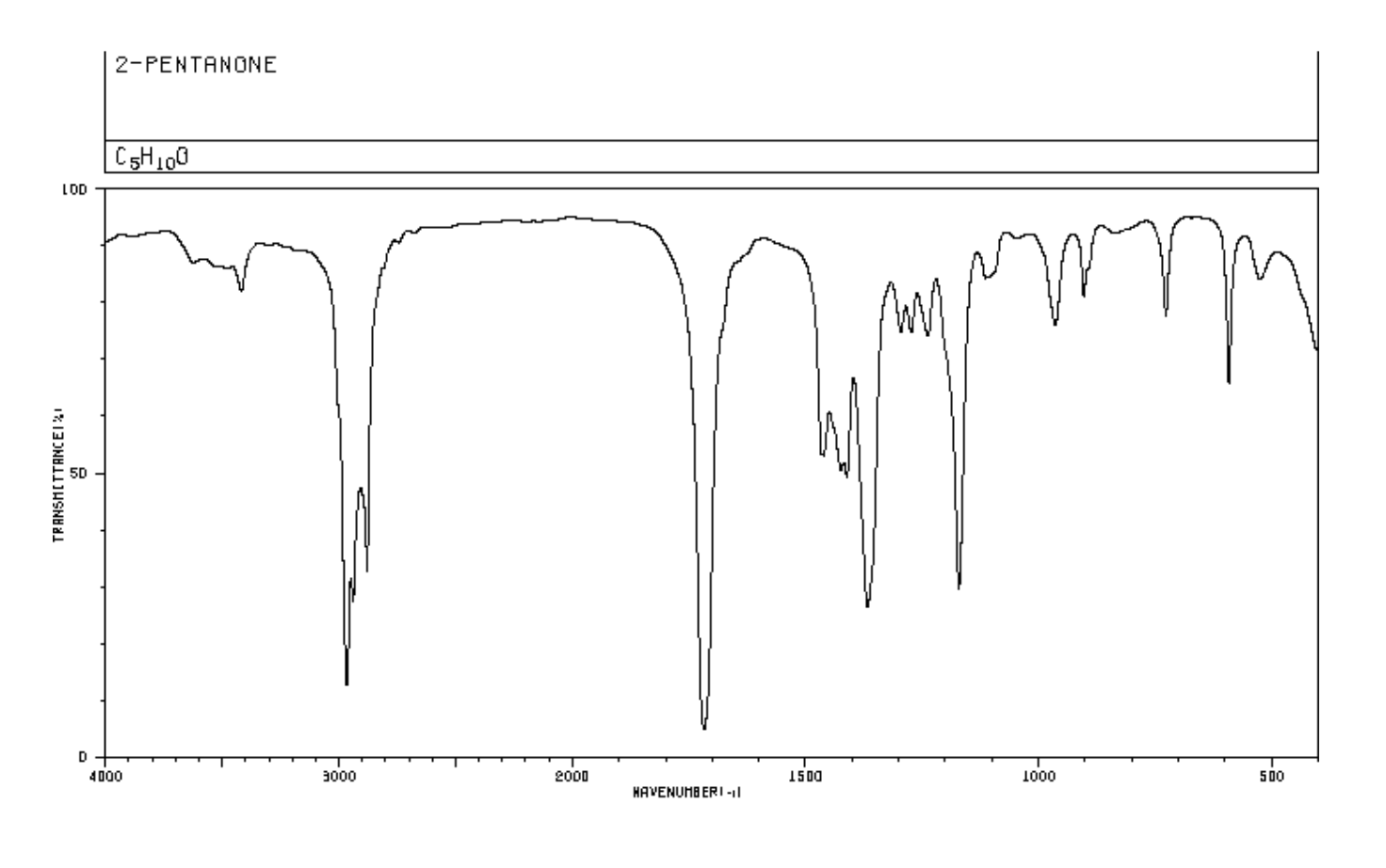
We have developed a Python-based deconvolution program, decon1d, which uses Bayesian information criteria (BIC) to objectively determine which model (number of peaks) would most likely produce the experimentally obtained data. One-dimensional (1D) 19F NMR spectra of proteins can be broad, irregular and complex, due to exchange of probe nuclei between distinct electrostatic environments and therefore cannot be deconvoluted and analyzed in an objective way using currently available software. Fluorine ( 19F) NMR has emerged as a useful tool for characterization of slow dynamics in 19F-labeled proteins. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
